Glossary
Glossary
You can find an even more comprehensive collection of information on the subject of research data management at https://forschungsdaten.info/themen/. You will also find many links in our RDM information collection.
Archiving / long-term storage
The archiving of research data is the unalterable and long-term storage of research data. To ensure the traceability of research results and good academic practice, research data must be stored for at least 10 years. Storage for more than 10 years is generally referred to as long-term archiving (LTA). In contrast to archiving for a limited period of time, additional requirements for permanent availability must be met for long-term archiving, such as the implementation of preservation strategies. The Open Archival Information System (OAIS) is a widely used standard for LTA.
It is particularly important in digital archiving that the data is stored unalterably and in a suitable format in order to ensure the preservation of the original data and legibility over the entire storage period. Other aspects that must be taken into account are the allocation of adequate access rights, adherence to deletion deadlines and description using metadata. The assignment of metadata is particularly important when archiving in a public repository so that the data is easier to find and reuse. However, this point should also not be neglected in the case of non-public archiving for reasons of traceability. If there is an obligation to delete data, which may be the case in medical research or personal data in particular, an archiving option should be selected where possible to comply with this deletion period, in which the data is automatically deleted after a specified period of time.
If research data is published in a public or institutional repository, it is automatically archived. If the data should or may not be made publicly accessible, the archiving service of your own institution can be used, if such a service exists. At the University of Oldenburg, only storage services are currently available for this purpose, which are difficult to use for archiving due to a lack of special functions. The introduction of a dedicated archiving system is currently being analysed/planned.
German Medicinal Products Act (AMG)
The German Medicinal Products Act is particularly relevant for research if a clinical trial as defined in Section 4 (23) is being conducted. In simple terms, these are all studies with medicinal products that are interventional in design, i.e. that contain randomisation or another form of determination that goes beyond the mere observation of medical practice or the intended purpose of the medicinal product. The KKS and the Medical Ethics Committee can advise you on this.
In the area of data management, the requirements of good clinical practice must be observed in particular. The services of the Research Data Management Service Centre are designed technically and organisationally in such a way that these requirements can be met. If our IT services are to be used in clinical trials, consultation with the Service Centre to define the processes and responsibilities is mandatory.
Audit Trail
In EDC systems, the audit trail refers to the recording or display/output of data changes. As a minimum, the time and user are recorded and saved and, in the case of changes/deletions, a comment/reason is sometimes also provided. This means that all changes can be traced (audited) at the level of individual data points.
The audit trail is an important component of quality management processes and is defined as a standard in Annex 11 of the EC GMP guidelines on computerised systems for GCP-compliant data processing.
Biobank / Biomaterial Bank (BMB)
A biobank is a collection of biomaterial, such as tissue and liquid samples, and the associated sample and subject data. Biobanks exist for the purpose of research, but an exact intended use is not determined at the time of sample collection. The samples and data are therefore usually stored for a long period of time, as specified in the consent documents, for no specific purpose or study.
The biomaterial is stored in the so-called sample bank, while the associated data is stored in databases. The associated data can be divided into different categories according to the TMF data protection guidelines [1]:
- Identifying data: This includes personal identifying data of the test subjects, such as name and date of birth. They usually remain at the place where the sample is taken (clinic/study centre) or are stored in a central patient list.
- Organisational data: The data contains information about the sample that is required to manage and categorise the sample. This includes, for example, the sample type, information on sample collection and storage.
- Analysis data: If samples are released to researchers, they can report their results back to the biobank. Like the sample itself, this analysis data can have the potential for re-identification, which must be taken into account when storing the data.
- Medical/research data: This includes socio-demographic data and clinical facts, e.g. diagnoses and findings, which are essential for the classification of the subject and thus for the research of the sample. The scope of this data, which is released to the researcher, depends on the scientific question.
For data protection reasons, the data in these categories should be available in pseudonymised form in physically and organisationally separate databases.
BBMRi-ERIC is a European biobank research data infrastructure that aims to facilitate access to biospecimens and promote progress in biomedical research. Germany is represented in this network by the German Biobank Network (GBN).
- Pommerening, K.; Drepper, J.; Helbing, K.; Ganslandt, T. (2014): Leitfaden zum Datenschutz in medizinischen Forschungsprojekten - Generische Lösungen der TMF 2.0, Schriftenreihe der TMF Band 11, MWV, Berlin.
Data Dictionary
A data dictionary is a structured documentation of the variable-related metadata of a research dataset. It clearly describes the variables contained in the data set as well as their meaning and characteristics, for example with regard to data types, permitted values or value ranges, coding, units of measurement, dependencies between variables and how to deal with missing values.
A data dictionary provides the technical and structural context of the data at the variable level and is a central prerequisite for correct analysis, the traceability and reproducibility of research results as well as the exchange and reuse of research data.
Ideally, a data dictionary should be created before data collection begins and maintained throughout the entire research process. It serves as a binding reference for data collection, data management, quality assurance, evaluation, archiving and publication of the data. Changes to the data dictionary after the start of data collection can affect data collection instruments, data quality and analysability.
In electronic data collection and data management systems, a data dictionary is often used for the systematic definition of a project data set. These systems usually provide functions that support the creation and editing of the data dictionary. For example, REDCap makes it possible to maintain the data dictionary via a graphical user interface (online designer) and via a standardised Excel/CSV import and export. Changes are checked by the system and implemented directly in the electronic data collection forms.
Data management plan (DMP)
A data management plan contains all relevant information about a project and the tasks that arise during the research data lifecycle. The use of a data management plan ensures that the individual data management steps are adequately described. In addition, the use of such a plan makes the handling of research data transparent, comprehensible and verifiable in terms of compliance. In some cases, the creation of one or, if necessary, several data management plans is expected by the funding body when applying for new projects, either as part of the application or within the first project period.
The Research Data Management Service Centre provides a sample template for the creation of a data management plan. Information on this can be found in our service area.
There are special software management plans for software that is developed as part of a research project.
Consent
Consent has different meanings depending on the area of law. In medical research, consent relevant under criminal law (e.g. Section 203 StGB and Section 228 StGB) and consent under data protection law in accordance with the GDPR are often relevant. This entry deals with consent under data protection law.
As a rule, special category personal data is processed in medical research. This category of data may only be processed in exceptional cases. The most common legitimisation ground here is consent (Art. 9 para. 2 lit. a GDPR). Consent under data protection law safeguards the informational self-determination of the person giving consent and at the same time enables the processing of personal data.
In addition, consent must be informed, specific and voluntary (Art. 4 no. 11 GDPR). Furthermore, the data may only be processed for the purposes specified in the consent (Art. 5 para. 1 lit. b GDPR). Further information and templates can be found on the pages of the Data Protection and Information Security Unit.
Consent management
Article 7 of the GDPR defines a number of conditions for handling consent. These include the obligation to provide proof of consent given (Art. 7 para. 1 GDPR) and the right to withdraw consent (Art. 7 para. 3 GDPR). Consent management is required to fulfil these conditions. Up to now, this has often been purely paper-based.
Digital consent management offers various options for supporting and improving consent processes:
- Obtaining consent: Consent can be recorded and signed digitally directly. Alternatively, paper-based consents can be scanned and recorded, meaning that the originals only need to be archived/stored.
- Consent management: Consents are stored digitally and are easily accessible for changes and reviews.
- Automated consent verification for data transfers: Consent management systems enable automated verification of consent for personal data
- Transparency: The recording and processing of consents and their metadata are standardised in consent management systems and enable the establishment of standardised processes. Traceability and standardisation simplify quality assurance.
- Increased data integrity: Digitisation and relocation to databases reduces the risk of data loss or irreparable damage to consent-related data, as backups can always be used to restore older data records.
- Data subject rights: Systematic and digital consent management helps to safeguard the data subject rights of study participants (Art. 15-21 GDPR), as the required information is easier to find and utilise.
Electronic Data Capture (EDC) / Electronic Case Report Form (eCRF)
The term Electronic Data Capture (EDC) refers to software systems that are used for the electronic recording or collection of (clinical) research data.
EDC systems are usually web-based and replace paper-based Case Report Forms (CRFs) with corresponding electronic representations (eCRFs) or paper-based surveys or self-documentation with suitable electronic variants. In addition to the input masks, EDC software usually also includes workflow support in the form of data entry checks, reporting/statistics modules, export modules and the like.
Other terms that are frequently used in this environment are Remote Data Entry (RDE), Clinical Data Management System (CDMS) and Survey Tool.
Further details can also be found in the glossary entries for the REDCap and SoSci Survey tools and on our service pages.
European Open Science Cloud (EOSC)
The European Open Science Cloud (EOSC) is a project of the European Commission to facilitate European scientists' access to scientific data, data processing platforms and data processing services.
The European Open Science Cloud (EOSC) ultimately aims to develop a 'Web of FAIR Data and services' for science in Europe upon which a wide range of value-added services can be built. These range from visualisation and analytics to long-term information preservation or the monitoring of the uptake of open science practices [1].
Among other things, the EU Node provides (free) IT services for research. This includes, for example
- File Sync and Share
- Interactive Notebooks (Jupyter)
- Large file transfer
- Virtual Machines
- Cloud Container Platform
- Bulk Data Transfer
The Resource Hub also contains well over 100 million individual entries in the categories:
- Publications
- Data
- Software
- Other Products
- Services
- Data Sources
- Training
- Interoperability Guidelines
- Tools
FAIR Data
The FAIR Data principles have long been included in guidelines for good academic practice. FAIR Data was explicitly published by the Force11 and is being further established, particularly as part of the GO FAIR initiative. In the German community, activities are currently being bundled within the framework of the National Research Data Infrastructure (NFDI).
The acronym FAIR is made up of the following terms:
F - Findable: In order to make data usable and reusable, it must be possible to find it. Machine-readable metadata that is suitable for searching is important for this.
A - Accessible: Found data must be accessible. This applies, among other things, to the communication protocols and, if necessary, authorisation management.
I - Interoperable: The metadata/data must be represented in a form that allows integration with other data and tools. This applies, for example, to the knowledge representation with regard to vocabularies, references, etc.
R - Reusable: To enable subsequent use of the data, suitable licences must be used, for example, the source and processing steps must be described(data provenance) and relevant domain/community standards must be observed.
Research data
On the one hand, research data is the result of scientific research activities and, on the other hand, represents an important basis for scientific work. In accordance with the diversity of scientific research, research data includes very different types of measured values, laboratory data, image or sound recordings, survey data, samples, etc. According to the guidelines on handling research data published by the German Research Foundation (DFG), methodological test procedures, e.g. questionnaires, software and simulations, should also be considered research data if they are a central result of scientific activity.
See also Lifecycle of research data, Metadata, FAIR Data.
Good Clinical Practice (GCP) and GxP
GxP is a collective term for regulatory guidelines in quality- and safety-critical research and application areas. In the field of research data management (RDM), GxP describes the framework for handling research data throughout the entire data life cycle , i.e. from collection, processing and analysis to archiving and, if necessary, submission to the authorities.
The GxP guidelines define requirements with regard to data quality, data integrity, documentation, traceability and access control. For RDM, this results in particular in the use of suitable and validated processes and electronic systems that provide audit-proof processing and storage of data. Ensuring data integrity is a central element of all GxP areas. This results in specific requirements for version control, audit trails, role and authorisation concepts as well as backup and archiving strategies.
GCP (Good Clinical Practice) maps the application of GxP quality standards to the area of clinical trials. The application of GCP is mandatory for the planning, conduct, documentation, evaluation and reporting of clinical trials on humans. Compliance with the GCP guidelines is intended to guarantee the safety and rights of the participants and ensure the credibility and accuracy of the data and results. The consideration of GCP in other clinical trials is recommended. The FDM Service Centre provides the EDC services secuTrial (validated for clinical trials, in co-operation with the KKS) and REDCap (for other clinical trials), among others. Both systems offer comprehensive support in the implementation of GCP requirements, for example through seamless audit trails, strict role and authorisation concepts, integrated query management and automated plausibility checks to ensure data quality.
Other GxP application areas include
- GLP (Good Laboratory Practice) - planning, implementation and monitoring of non-clinical health and environmental safety tests in a laboratory-like environment
- GMP (Good Manufacturing Practice) - production of medicinal products and active ingredients
Good academic practice (GWP)
Scientific honesty and compliance with the principles of good academic practice are indispensable prerequisites for scientific work that aims to gain knowledge for the benefit of society.
To this end, the University of Oldenburg has adopted the "Regulations on the Principles for Safeguarding Good Academic Practice at the University of Oldenburg". The regulations and further information can be found on the pages of the School and the Commission of the Senate.
The current version of the "Guidelines for Safeguarding Good Academic Practice" can be found on the DFG website.
The Research Data Management Service Centre can support you with the implementation of the GWP, particularly in the following areas:
- Guideline 7: Cross-phase quality assurance
Advice on subject-specific standards and established methods in the areas of research data management, medical informatics, data protection, information security, etc.
Quality assurance through the design of state-of-the-art data management processes in the collection, processing and analysis of research data as well as the selection, use, development and programming of research software, where applicable. - Guideline 8: Actors, responsibilities and roles
If IT services are used that are administered by the service centre, we will maintain the user roles for you. The associated rights and responsibilities are therefore transparent at all times during the research project. - Guideline 10: Legal and ethical framework conditions, rights of use
The Service Agency can support you with regard to legal admission, licences of use and the technical implementation of making research data and results accessible. - Guideline 11: Methods and standards
The Service Centre can provide you with methodological support, particularly in the areas of research data management, medical informatics, data protection and information security. In line with the guideline, you can utilise these skills, for example, when using software and collecting research data. - Guideline 12: Documentation
By using data management plans and documentation tools, you provide the necessary information about the research data and the processing steps in accordance with the guideline. - Guideline 13: Providing public access to research results
In this area, the Service Centre is working with the university's central institutions on solutions to better support the FAIR publication of research data. - Guideline 16: Confidentiality and neutrality in reviews and consultations
The obligation of confidentiality naturally also applies to consultations by the Research Data Management Service Centre. - Guideline 17: Archiving
We will be happy to support you in implementing the retention period, which is generally 10 years.
Identity management (patient list, anonymisation, pseudonymisation, trustee office)
Identity management is used in medical research to process medical data (MDAT) in accordance with the principles of the GDPR (in particular Article 5(1)(c) and (e)) in a data protection-compliant manner. Identity management is necessary when anonymous data cannot be used in research, such as when merging data from different sources or in the case of longitudinal data. This process is also known as record linkage. The central tasks of identity management include maintaining patient lists and pseudonymisation.
In the document "Practical guide to anonymisation/pseudonymisation" [3] you will find further definitions, legal classifications, implementation recommendations, practical examples, checklists, etc.
Patient list
In a patient list, persons are recorded with their identifying personal data (IDAT) and assigned a non-speaking identifier (PID) (in practice often also called study patient number or similar). The PID serves as an identifier instead of the IDAT when transferring treatment or Study details to the research context.
Identity management can be organised centrally or decentrally. Patient lists can be kept locally within a research project in the interests of data minimisation. For larger research networks with data from different sources, it is often advisable to have patient lists and pseudonymisation operated by a central, independent and trustworthy body.
Anonymisation
Anonymisation occurs when the personal reference of data is removed in such a way that it cannot be restored or can only be restored with a disproportionate amount of time, cost and manpower. [2]
Pseudonymisation
Article 25 of the GDPR requires pseudonymisation when processing personally identifiable data. According to [1], different classes of pseudonyms are differentiated according to the group of people who become aware of them. Multi-level pseudonymisation can be used to securely decouple the context of care, a research database and any extracts derived from it.
Pseudonymisation service
If medical data from different sources is to be merged, this is done using pseudonymisation. A permanent pseudonym (PSN) is assigned to a PID from identity management. The associated MDAT is passed through a pseudonymisation service or encrypted.
In this context, a pseudonymisation service is a software solution that enables the automatic pseudonymisation of data. These systems usually offer the automatic creation, storage and management of pseudonyms and the assignment of original values to pseudonyms. There are both simple desktop applications for small local research projects and complex client-server architectures for distributed research networks.
Trust centre / Trusted authority
In medical research, a trust centre serves as an independent institution for the separation of personal identifying data and medical data. In the glossary entry Repository you will find information on the term "trust centre" in a non-medical context, which is used in the Federal Government's data strategy.
Data can be made available to researchers through a trust centre without a direct personal reference in such a way that they can still be linked via pseudonyms. Trust centres can be operated locally, distributed or centrally. In addition to maintaining a patient list and pseudonymising or anonymising data, a trust centre can also manage consent forms from patients and test subjects.
At the beginning of 2022, the UOL decided to establish the independent (data) trust centre of the University of Oldenburg.
For many of the tasks under the umbrella term identity management, software solutions can be offered to facilitate implementation. For example, a pseudonymisation service could take over the systematic and automatic pseudonymisation of data.
You can find the FDM Service Centre's offers in this regard in our service area. Further offers, in particular for technical implementation, can be found at the Fiduciary Agency. The main contact for technical implementation is the Fiduciary Agency.
- Pommerening, K.; Drepper, J.; Helbing, K.; Ganslandt, T. (2014): Guidelines on data protection in medical research projects - Generic solutions of TMF 2.0, TMF publication series volume 11, MWV, Berlin.
- The Federal Commissioner for Data Protection and Freedom of Information (2020): Public consultation procedure of the Federal Commissioner for Data Protection and Freedom of Information on the topic: Anonymisation under the GDPR with special consideration of the telecommunications industry, Bonn.
- Practical guide to anonymisation/pseudonymisation from the GMDS working group "Data protection and IT security in the healthcare sector" and the BvD (as of 27.01.2024) https://gesundheitsdatenschutz.org/html/pseudonymisierung_anonymisierung.php
Investigator Initiated Trials (IIT) / science-initiated (clinical) studies
Investigator Initiated Trials (IITs) are clinical studies or biomedical research projects for which the responsibility (i.e. sponsorship according to the German Medicines Act) and the study management lies with university institutions. In contrast, approval studies in the pharmaceutical or medical device sector are usually conducted by the (pharmaceutical) industry in collaboration with contract research organisations (CROs).
Typical examples of IITs are therapy comparisons or therapy optimisation studies, for which there is often no commercial interest but a high scientific interest.
IITs are often financed by public funding programmes such as the BMBF or DFG , by foundations and sometimes also by industry.
Life cycle of research data
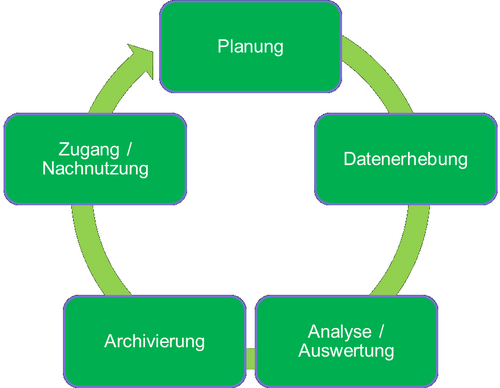
The life cycle of research data extends from the planning of a research project through the generation and analysis of research data to the archiving and utilisation of research data. Various generic phases can be identified in the life cycle of research data:
Planning: the starting point of a research project is always a research question. In evidence-based research, data is required to answer this question. This involves a variety of tasks:
- Researching existing data and data access options
- Clarification of data licence issues
- Planning data collection
- Planning research data management, including
- Planning data access
- Planning data publication
- Planning data archiving
When conducting research with research data, it is advisable to practise systematic research data management, to develop a data management plan during the planning phase and to maintain it continuously during the research project.
- Data collection: Data collection includes tasks ranging from the pure acquisition of existing data to the collection of new data (e.g. through experiments or surveys). Depending on the research project, software tools can be used to collect new data. These can be EDC solutions such as REDCap or survey servers such as SoSci Survey can be used. Furthermore, data collection also includes the collection of metadata to describe the data and the context of the survey and the management and storing data.
- Analysis/evaluation: In order to analyse and evaluate the research data, the research data must be checked, validated and, if necessary, cleansed. This represents an interface between data collection and analysis. The research data can then be analysed and interpreted. The findings obtained are often available in the form of analysis data and are also research data.
- Archiving: After analysis/evaluation and, if applicable, publication of scientific results, the research data must be archived. Archiving should already be prepared in the planning phase and documented in a data management plan documented in a data management plan. Archiving tasks include the selection of an archive and/or archiving method, selection of a suitable storage format and corresponding documentation. Furthermore, archiving can also include data preparation tasks, such as anonymisation of data records.
- Access/reuse: Access to and subsequent use of research data are based on the FAIR principles. Making research data accessible for subsequent scientific use includes the publication of research data in an accessible research data repository and the granting of licences to use the research data. At the beginning of this phase, the legal question of the form in which the research data may be made accessible at all must also be clarified. Data protection hurdles often need to be considered here, especially in the context of medical research.
Licences and rights of use
Licences and usage rights are important in the context of research for the (re-)use of software, but also for data and texts. This is relevant both from the perspective of the author/provider and for users. The subject area is very complex and extensive - special cases may require individual legal advice. Please also refer to our general FAQ article on the topic of legal issues. The aim here is merely to provide a brief overview and refer you to general collections of information.
A good introduction to legal issues relating to publication can be found at: https://forschungsdaten.info/themen/rechte-und-pflichten/recht-und-forschungsdaten-ein-ueberblick/
Before the publication of research data and software, it must be clarified whether or not a licence is necessary and appropriate. Copyright law protects works that reach a "level of creation", and only such works require licences that allow other users to use these works. There are differences between software, (measurement) data, databases and texts/images/art objects. You can use this flowchart to check the most important conditions for publishing research data yourself.
When using software, the applicable (possibly also commercial) licence conditions must be observed. When publishing (your own) software, open source licences should be used wherever possible. A comprehensive overview of open source licences can be found at: https://opensource.org/licenses. A widely used licence is, for example, the GNU General Public License (GPL). When modifying/redistributing third-party source code, the choice may be limited, as the respective conditions must be observed (e.g. "Copyleft": derived works must be licensed under the same conditions).
Data - such as measurement results in tabular form - are generally not protected by copyright. The necessary individuality and originality are often not given. However, special rules apply to databases (systematically or methodically organised collections of works or data whose procurement, review or presentation requires a substantial investment in terms of type or scope). For data publications, for example, https://opendatacommons.org/licenses/ comes into question.
Creative Commons licences, for example, can be used for general works protected by copyright.
Medical terminology
In medical research and care, there is a high demand for semantic interoperability for the exchange of medical data. Semantic interoperability is intended to ensure the preservation of meaning in the exchange of information between humans and machines. Examples of medical terminologies are
- ICD: The International Statistical Classification of Diseases and Related Health Problems (ICD) is an important globally recognised classification system for medical diagnoses. The German version of the currently valid ICD version 10: https://www.bfarm.de/DE/Kodiersysteme/Klassifikationen/ICD/ICD-10-GM/_node.html
- LOINC: Logical Observation Identifiers Names and Codes (LOINC) is an international, primarily English-language system for the unique coding of medical examinations, in particular laboratory examinations: loinc.org/
- SNOMED CT: Systematised Nomenclature of Medicine Clinical Terms (SNOMED CT) is currently one of the most comprehensive and important medical terminologies worldwide: www.snomed.org/
Medical devices / Medical Device Law Implementation Act (MPDG) / Medical Device Regulation (MDR)
Medical devices are products with a medical purpose that are intended by the manufacturer for use in humans.
These include implants, products for injection, infusion, transfusion and dialysis, human medical instruments, software, catheters, pacemakers, dental products, dressings, visual aids, X-ray equipment, condoms, medical instruments, laboratory diagnostics, products for contraception and in vitro diagnostics.
In vitro diagnostics include reagents, reagent products, kits, sample containers, devices and other products intended for the in vitro examination of samples from the human body.
Medical devices are also products that contain or are coated with a substance or preparations of substances which, when used separately, are considered to be medicinal products or components of medicinal products (including plasma derivatives) and may have an effect on the human body in addition to the functions of the product.
In contrast to medicinal products, which have a pharmacological, immunological or metabolic effect, the main intended effect of medical devices is primarily achieved by physical means [1].
The purpose of the regulations (in the research context) is, among other things, to ensure that the data obtained in clinical trials is reliable and robust and that the safety of trial participants is protected.
The legal basis for this includes the Medical Device Regulation (MDR) and the Medical Device Law Implementation Act (MPDG).
Please contact the KKS regarding the regulatory categorisation of studies and the resulting requirements.
Metadata
Metadata is structured data that contains information about the characteristics of other data. In the context of research data , information about the creation/acquisition and processing of the data is among the most important metadata. This ranges from general project-related information (research area, project leader, funding, ...) to detailed technical information for imaging or (laboratory) analytical methods.
Comprehensive general information on the topic of research data with many sources and references can be found in the article Metadata and metadata standards.
The publication 5 Steps to Better Metadata and the decision tree Difficult Decision - How do I deal with metadata? also offer a good introduction with practical tips.
Metadata is an important basis for FAIR Data.
National Research Data Infrastructure (NFDI)
The National Research Data Infrastructure is a funding initiative coordinated by the DFG to develop a reliable and sustainable portfolio of services for generic and subject-specific research data management needs throughout Germany.
The National Research Data Infrastructure (NFDI) is intended to systematically develop, sustainably secure and make accessible the data holdings of science and research and to network them (inter)nationally. It will be established as a networked structure of consortia acting on their own initiative in a process driven by the scientific community.
The aims of funding consortia are
- Establishing rules for the standardised handling of data in close cooperation with the respective specialist community
- Development of cross-disciplinary metadata standards
- Development of reliable and interoperable measures for data management and a range of services tailored to the requirements of the specialist community
- Increasing the reusability of existing data, including across disciplinary boundaries
- Connecting and networking with partners in foreign scientific systems that have expertise in the field of research data management.
- Collaboration in the development and establishment of generic, cross-consortia services and standards for research data management [1]
In the medical field, NFDI4Health, which deals with health data, and Base4NFDI, which integrates and establishes basic RDM services, are particularly important.
At European level, the NFDI co-operates with the EOSC. Together with colleagues from the BIS, the Department for Research and Technology Transfer and the IT services, the Research Data Management Service Centre follows developments in this area and provides information on services and solutions.
Persistent identifiers (PIDs)
Persistent identifiers (PIDs) are permanent, digital identifiers that enable the long-term and unique referencing of digital objects, persons and organisations. Their use contributes significantly to the citability, findability and reusability of research data.
The DOI (Digital Object Identifier) and the URN (Uniform Resource Name) are assigned to digital objects and ensure that objects (e.g. data and publications) can be found reliably even if the storage location or URL changes.
Researchers can register with ORCID (Open Researcher and Contributor ID) to uniquely identify individuals. Published data can be added to the ORCID profile automatically or manually from various sources so that the research results can be collected centrally and clearly linked to the respective person.
Organisations are identified via the ROR ID (Research Organisation Registry ID). The University of Oldenburg recommends the use of ROR-ID and ORCID in its affiliation guidelines.
Further information on persistent identifiers is available at forschungsdaten.info.
Personal data
According to Article 4 of the General Data Protection Regulation (GDPR), personal data is any information relating to an identified or identifiable person.
Health data in the form of individual details (e.g. a table with one line per person) is therefore generally personal data. Even if no directly identifying characteristic (e.g. the name) or a pseudonym (e.g. a case number) is included, a person can still be identifiable.
A legal basis (such as the consent of the data subject) is required for the processing of personal data. This is particularly important if data collected for other reasons (such as medical care) is to be used for research and transferred to third parties.
In accordance with Article 9 of the GDPR, medical data are regularly so-called "special categories of personal data". In particular, this includes genetic data, biometric data and health data. These types of data are considered particularly sensitive and therefore require additional measures.
The Research Data Management Service Centre will be happy to advise you on special aspects of handling medical research data. The topic of identity management, for example, is also relevant in this context.
Further details and advice can also be found on the pages of the Data Protection and Information Security Unit.
REDCap
REDCap is a browser-based EDC software for building and managing clinical and translational research databases. Data collection is convenient and simple via a user-friendly interface with the help of online surveys or eCRFs .
The collected data can be exported as CSV or XML or directly in the format for various statistical tools (SPSS, SAS, R, Stata). REDCap also offers offline data collection via app, an audit trail, the retrieval of reports and statistics, extensibility through plugins and much more.
A detailed list of the functions included can be found on the official website: projectredcap.org/software/
It should be noted that the software is not validated for studies subject to the German Drug Law (AMG) or the German Medical Devices Act (MPG).
REDCap was implemented at Vanderbilt University in 2004 and has been continuously developed by the REDCap consortium ever since.
Information on our REDCap service can be found in our service area.
Repository
In the research context, a repository is a storage location for research data (e.g. raw data, methodological information, publications). It can be used to make research data accessible to the public and to store it for a limited period of time or to archive it permanently. Publishing research data in a repository is in line with good academic practice (GWP) and the FAIR Data principles and makes it much easier for other researchers to find and re-use the data. In addition to general repositories, there are subject repositories that specialise in a particular discipline and the specifics of the respective data, and institutional repositories that are only accessible to members of the respective institution. At the University of Oldenburg, such a repository called DARE is provided by BIS.
If the research data or publications are entered in the repository, they are generally assigned a persistent identifier (PID). This ensures that the data and publications can always be referenced and cited even if the storage location changes. The DOI (Digital Object Identifier) and the URN (Uniform Resource Name) are frequently used for this purpose. To ensure that the data can be found and reused, the data must be adequately described by metadata . The metadata also contains the licences that specify the conditions under which the data may be used. The repository operator may also have special requirements here. Before inclusion in the repository, particular attention should be paid to legal aspects (copyright, secondary publication, data protection; see also FAQ). In some cases, this is checked by the operator, as is data curation (quality control of content, metadata annotation, data format, etc.).
Research data repositories can be found via research data repository registers, such as the Registry of research data repositories [1]. It is also often helpful to look at outstanding publications in your own field of research to identify relevant specialised repositories.
In the context of the provision of FAIR data by public repositories, the political development in this area is worth mentioning. On 27 January 2021, the Federal Government's data strategy was adopted with around 240 measures, which aims to "significantly increase the provision and use of data by individuals and institutions [...], prevent the creation of new data monopolies, ensure fair participation and at the same time consistently counter data misuse" [2]. This refers to data trusts, which are intended as neutral bodies for the administration and provision of data and thus fulfil the functions of a repository to a certain extent. Further information and the full version of the data strategy can be found on the German government's website [3]. The European Commission has also spoken out in favour of promoting common European data spaces and the resulting data pools in strategic economic sectors and areas of public interest [4]. The Federal Government also sends out strong signals in the 2023 update of the data strategy "Progress through data use" - for example with the chapter headings "More data", "Better data" and "Data use and data culture"[5].
The full statement can be found on the RfII website.
- www.forschungsdaten.info/themen/veroeffentlichen-und-archivieren/repositorien/
- The Federal Government (2019): Key points of a data strategy, p. 1.
- www.bundesregierung.de/breg-de/themen/digitalisierung/datenstrategie-beschlossen-1842786
- European Commission (2020): A European data strategy. Communication from the Commission to the European Parliament, the Council, the European Economic and Social Committee and the Committee of the Regions, Brussels, p. 25.
- The Federal Government: Progress through data use (as at August 2023)
Research Software Marketplace
A research software marketplace is an online platform that provides access to software tools and services that are specifically suitable for use in research. This type of platform can also be found under other names, such as research software directory or software catalogue. With the help of filter and search options, researchers can search the platform for suitable tools for their discipline and use case - e.g. data collection, data analysis, visualisation or modelling - and thus reuse existing software instead of having to carry out development work themselves.
There are different types of research software marketplaces that vary in scope and focus. Some platforms focus on a specific discipline, while others offer general and specialised tools for different disciplines. They also differ in terms of whether only open software or commercial and fee-based tools are included. The free products are usually software that has to be installed locally by the researchers themselves. As a rule, open software can be reused, i.e. it can be further developed and customised to suit individual requirements. Licence terms, good academic practice and, where applicable, usage rights must be observed. The DFG also has information on how to use research software on its website.
The EOSC 's Resource Hub (contains e.g. software (publications), tools and (IT) services ) and the nfdi.software marketplace, which is still under construction, are two larger portals that contain open and free software. The TMF's Health Research ToolPool specialises in medical research. In addition to open tools, it also lists commercial tools and offers further services, such as counselling services and working materials.
Software as a Service (SaaS)
Software as a service is a sub-sector of cloud computing. According to the German Federal Office for Information Security (BSI), it is defined as follows:
Cloud computing refers to the dynamic provision, use and settlement of IT services via a network in line with demand. These services are offered and utilised exclusively via defined technical interfaces and protocols. The range of services offered as part of cloud computing covers the entire spectrum of information technology and includes infrastructure (e.g. computing power, storage space), platforms and software. [1]
SaaS is concerned with the provision of software as a service for the user. This means that the user no longer has to deal with the operation and basic administration of IT systems and the respective application. SaaS enables the user to focus on the specialised use of the respective software solution for a project.
Software Management Plan / Software Engineering Guideline / Development of FAIR research software
If software is developed as part of a research project, a software management plan should be drawn up (analogous to a data management plan). Software can vary in scope and complexity, from short shell scripts and extensive analysis programmes to complex software development projects. There are different recommendations depending on the complexity and, in particular, the intended purpose/application class.
A document with extensive recommendations and checklists in both German and English under a CC-BY licence is available from the DLR Software Engineering Initiative [1].
The Software Sustainability Institute [2] also has a recommended overview page with linked templates and information.
In October 2024, the DFG published a paper on "Dealing with research software in the DFG's funding activities" [3]. It provides a well-structured overview of the topic and refers to other international initiatives and publications, e.g. FAIR4RS [4] and the ReSA Software Policies [5]. It also contains practical tips for project proposals and recommendations for the further development of the structural framework.
SoSci Survey
SoSci Survey is a professional browser-based tool for developing and conducting online surveys. The basis for SoSci Survey was created in 2003 at the Institute for Communication Science and Media Research at LMU Munich. Since then, SoSci Survey has been continuously developed by SoSci Survey GmbH.
SoSci Survey enables the creation of sophisticated online questionnaires by allowing images, audio and video files to be integrated into the questionnaire and providing over 30 question types for flexible design. The collected data can be exported in various formats such as CSV, SPSS, Stata, GNU R or SQL for further processing. Further information can be found on the official homepage: www.soscisurvey.de/
The use of SoSci Survey is free of charge for non-commercial research. The University of Oldenburg provides its own survey server with SoSci Survey for this purpose. The Research Data Management Service Centre offers the opportunity to use SoSci Survey at the University of Oldenburg for your own online surveys.
Information on our SoSci Survey service can be found in our service area.
Standard Operating Procedure (SOP)
Standard Operating Procedures (SOPs) are bindingly documented workflows or process descriptions. SOPs are an important component of a GCP-compliant quality management system.
Accordingly, an SOP system also includes a role model and a training concept. An SOP should precisely define its purpose and area of application as well as describe the relevant regulatory requirements and the work steps to be carried out, including the documentation to be created.
SOPs are often supplemented by further work instructions in which, for example, system-specific work steps are described in more detail compared to the more general requirements of an SOP.
TMF e. V.
The TMF - Technology and Methods Platform for Networked Medical Research e.V. (TMF for short) is the umbrella organisation for collaborative medical research in Germany. It is the platform for interdisciplinary exchange and cross-project and cross-site cooperation in order to jointly identify and solve the organisational, legal-ethical and technological problems of modern medical research. The solutions range from expert opinions, generic concepts and IT applications to checklists, guidelines, training and counselling services. The TMF makes these solutions freely and publicly available. [1]
The School of Medicine is a member of the TMF and would like to make the services available to members of the UOL. Employees of the service centre regularly take part in TMF working group meetings and are happy to answer your questions or to establish contacts / co-operations.
